Curran Jude
PointNu-Net: Simultaneous Multi-tissue Histology Nuclei Segmentation and Classification in the Clinical Wild
Nov 01, 2021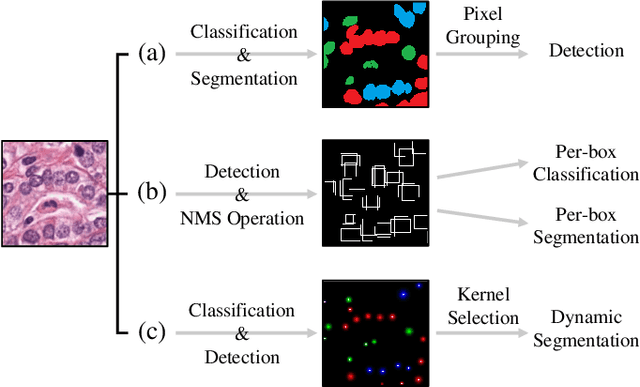
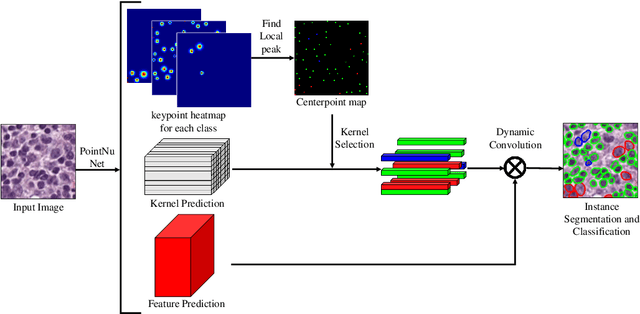
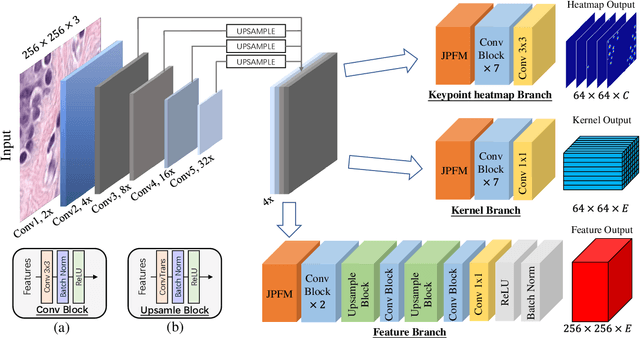
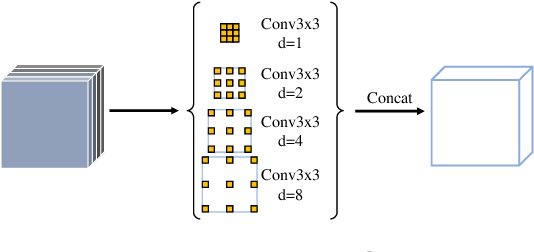
Abstract:Automatic nuclei segmentation and classification plays a vital role in digital pathology. However, previous works are mostly built on data with limited diversity and small sizes, making the results questionable or misleading in actual downstream tasks. In this paper, we aim to build a reliable and robust method capable of dealing with data from the 'the clinical wild'. Specifically, we study and design a new method to simultaneously detect, segment, and classify nuclei from Haematoxylin and Eosin (H&E) stained histopathology data, and evaluate our approach using the recent largest dataset: PanNuke. We address the detection and classification of each nuclei as a novel semantic keypoint estimation problem to determine the center point of each nuclei. Next, the corresponding class-agnostic masks for nuclei center points are obtained using dynamic instance segmentation. By decoupling two simultaneous challenging tasks, our method can benefit from class-aware detection and class-agnostic segmentation, thus leading to a significant performance boost. We demonstrate the superior performance of our proposed approach for nuclei segmentation and classification across 19 different tissue types, delivering new benchmark results.
AD-GAN: End-to-end Unsupervised Nuclei Segmentation with Aligned Disentangling Training
Jul 23, 2021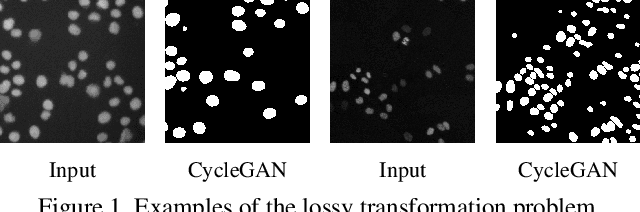
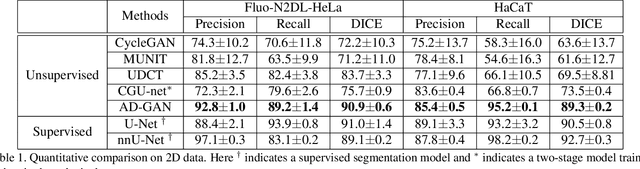
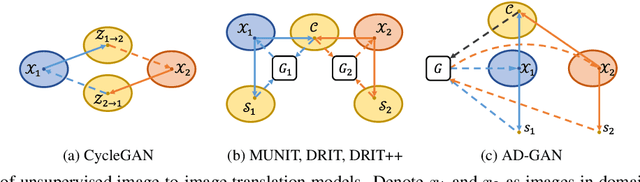
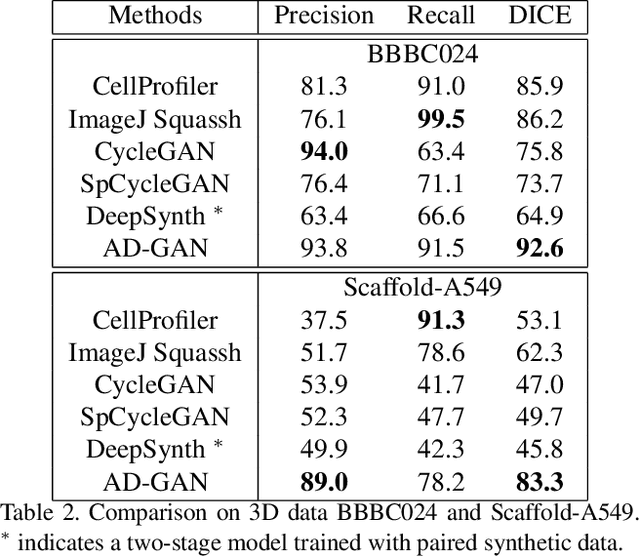
Abstract:We consider unsupervised cell nuclei segmentation in this paper. Exploiting the recently-proposed unpaired image-to-image translation between cell nuclei images and randomly synthetic masks, existing approaches, e.g., CycleGAN, have achieved encouraging results. However, these methods usually take a two-stage pipeline and fail to learn end-to-end in cell nuclei images. More seriously, they could lead to the lossy transformation problem, i.e., the content inconsistency between the original images and the corresponding segmentation output. To address these limitations, we propose a novel end-to-end unsupervised framework called Aligned Disentangling Generative Adversarial Network (AD-GAN). Distinctively, AD-GAN introduces representation disentanglement to separate content representation (the underling spatial structure) from style representation (the rendering of the structure). With this framework, spatial structure can be preserved explicitly, enabling a significant reduction of macro-level lossy transformation. We also propose a novel training algorithm able to align the disentangled content in the latent space to reduce micro-level lossy transformation. Evaluations on real-world 2D and 3D datasets show that AD-GAN substantially outperforms the other comparison methods and the professional software both quantitatively and qualitatively. Specifically, the proposed AD-GAN leads to significant improvement over the current best unsupervised methods by an average 17.8% relatively (w.r.t. the metric DICE) on four cell nuclei datasets. As an unsupervised method, AD-GAN even performs competitive with the best supervised models, taking a further leap towards end-to-end unsupervised nuclei segmentation.
 Add to Chrome
Add to Chrome Add to Firefox
Add to Firefox Add to Edge
Add to Edge